|
|
Vol.
32 No. 3
May-June 2010
Metallomics: Guidelines for Terminology and Critical Evaluation of Analytical Chemistry Approaches (IUPAC Technical Report)
Ryszard Lobinski, et al.
Pure and Applied Chemistry, 2010
Vol. 82, No. 2, pp. 493–504
The knowledge of the complete genetic blueprint of an increasing number of organisms has resulted in efforts aimed at the global analysis and functional study of a particular class of components of a living organism and the emergence of different “-omics.” The concepts of genomics (the study of genes and their function) and proteomics (the study of the set of proteins produced by an organism, their localization, structure, stability, and interaction) have become part of the everyday language of life sciences. Metal ions are a vital component of the chemistry of life. One-third of all proteins is believed to require a metal cofactor, such as copper, iron, zinc, or molybdenum, delivered as a simple or complex ion or a metal-containing compound (e.g., methylcobalamin). The intracellular concentration of several metals, their distribution among the various cell compartments, and their incorporation in metallo proteins are tightly controlled. The understanding of mechanisms by which a metal is sensed, stored, or incorporated as a cofactor requires, in addition to the identification of metalloproteins, the characterization of the pool of nonprotein molecules (products of enzymatic or biochemical reactions) interacting with metal ions or of metabolites of exogenous metallocompounds, such as metallodrugs. A systematic approach to the study of metal content, speciation, localization, and use in biological systems is becoming increasingly important.
The term “metallome” was coined by Williams who referred to it as an element distribution, equilibrium concentrations of free metal ions, or as a free element content in a cellular compartment, cell, or organism. The latter would therefore be characterized not only by its genome or proteome but also by the metallome, their inorganic complement. The meaning of the term “metallome” was then proposed to be extended to the entirety of metal and metalloid species present in a cell or tissue type.
The characterization of the pool of metal-containing species in living organisms and of their interactions with the genome, transcriptome, proteome, and metabolome requires dedicated analytical approaches to in vivo detection, localization, identification, and quantification, in vitro functional analysis, and “in silico” prediction using bioinformatics. The term “metallomics” was coined by Haraguchi to denote the ensemble of research activities related to metals of biological interest (see figure).
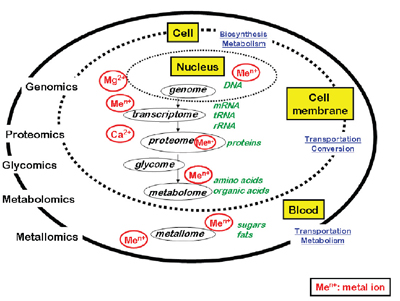 |
Simplified model of biological system and related -omics sciences. The outer area surrounded with the continuous line is showing, e.g., an organ or whole body, and the inner area surrounded with the dotted line is showing a biological cell. Biological fluid, e.g., blood, is circulating in the intermediate area. The Mg2+ and Ca2+ ions are given as examples because of their large affinities with DNA and proteins, respectively, in the biological cell. |
Metallomics has been the topic of a number of reviews, special issues of edited journals and of a Royal Society of Chemistry journal dedicated to the field, Metallomics. The terms “metallome” and “metallomics” have been used in different contexts. In addition, a number of related definitions proliferated, such as, for example, ionomics, heteroatom-tagged proteomics, or elementomics. This report attempts to define the terms “metallome” and “metallomics,” critically evaluates the available relevant analytical methodology, and highlights the concerned disciplines and research areas.
Note: Figure and intro reproduced from PAC; see PAC for full text and references.
http://dx.doi.org/10.1351/PAC-REP-09-03-04
Page
last modified 11 May 2010.
Copyright © 2003-2010 International Union of Pure and Applied Chemistry.
Questions regarding the website, please contact [email protected] |